Research Projects - Other
The nature of modern science is that it is ever-changing, energetically crossing boundaries heretofore defined by traditional areas of inquiry. Research at the Theoretical and Computational Biophysics group reflects this dynamic, with studies employing theoretical perspectives and methodological approaches or addressing topics that don't fall easily into one of the above categories. Included in this broad category are studies of a four-way DNA junction, the nuclear pore complex, gas transport in hydrogenase that may provide a source of renewable fuel, and other topics.
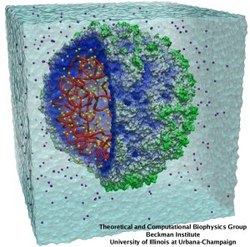
image size:
150.4KB
made with VMD
Viruses, the cause of many diseases, are the smallest natural organisms known. They are extremely primitive and parasitic such that biologists refer to them as "particles", rather than organisms. Viruses contain in a protein shell, the capsid, their own building plan, the genome, in the form of DNA or RNA. Viruses hijack a biological cell and make it produce from one virus many new ones. Viruses have evolved elaborate mechanisms to infect host cells, to to produce and assemble their own components, and to leave the host cell when it bursts from viral overcrowding. Because of their simplicity and small size, computational biologists selected a virus for their first attempt to reverse engineer in a computer program, NAMD, an entire life form, choosing one of the tiniest viruses for this purpose, the satellite tobacco mosaic virus. As described in a recent report, the researchers simulated the virus in a small drop of salt water, altogether involving over a million atoms. This provided an unprecedented view into the dynamics of the virus for a very brief time, revealing nevertheless the key physical properties of the viral particle as well as providing crucial information on its assembly. It may take still a long time to simulate a dog wagging its tail in the computer, but a big first step has been taken to simulate living organisms. Naturally, this step will assist modern medicine (more on our satellite tobacco mosaic virus web page).
Papers
Multilevel summation with B-spline interpolation for pairwise interactions in molecular dynamics simulations. David J. Hardy, Matthew A. Wolff, Jianlin Xia, Klaus Schulten, and Robert D. Skeel. Journal of Chemical Physics, 144:114112, 2016. (16 pages).
Multilevel summation method for electrostatic force evaluation. David J. Hardy, Zhe Wu, James C. Phillips, John E. Stone, Robert D. Skeel, and Klaus Schulten. Journal of Chemical Theory and Computation, 11:766-779, 2015.
Mature HIV-1 capsid structure by cryo-electron microscopy and all-atom molecular dynamics. Gongpu Zhao, Juan R. Perilla, Ernest L. Yufenyuy, Xin Meng, Bo Chen, Jiying Ning, Jinwoo Ahn, Angela M. Gronenborn, Klaus Schulten, Christopher Aiken, and Peijun Zhang. Nature, 497:643-646, 2013.
A computational kinetic model of diffusion for molecular systems. Ivan Teo and Klaus Schulten. Journal of Chemical Physics, 139:121929, 2013. (15 pages).
Effects of cytosine hydroxymethylation on DNA strand separation. Philip M.D. Severin, Xueqing Zou, Klaus Schulten, and Hermann E. Gaub. Biophysical Journal, 104:208-215, 2013.
DNA target sequence identification mechanism for dimer-active protein complexes. Markita P. Landry, Xueqing Zou, Lei Wang, Wai Mun Huang, Klaus Schulten, and Yann R. Chemla. Nucleic Acids Research, 41:2416-2427, 2013.
A computational kinetic model of diffusion for molecular systems. Ivan Teo and Klaus Schulten. Journal of Chemical Physics, 139:121929, 2013. (15 pages).
Decrypting cryptochrome: Revealing the molecular identity of the photoactivation reaction. Ilia A. Solov'yov, Tatiana Domratcheva, Abdul R. M. Shahi, and Klaus Schulten. Journal of the American Chemical Society, 134:18046-18052, 2012.
Further optimization of a hybrid united-atom and coarse-grained force field for folding simulations: Improved backbone hydration and interactions between charged side chains. Wei Han and Klaus Schulten. Journal of Chemical Theory and Computation, 8:4413-4424, 2012.
Molecular basis of drug resistance in A/H1N1 virus. Ariela Vergara-Jaque, Horacio Poblete, Eric Lee, Klaus Schulten, Fernando González-Nilo, and Christophe Chipot. Journal of Chemical Information and Modeling, 52:2650-2656, 2012.
Unique sugar-binding site mediates the distinct anti-influenza activity of pig surfactant protein D. Martin van Eijk, Michael J. Rynkiewicz, Mitchell R. White, Kevan L. Hartshorn, Xueqing Zou, Klaus Schulten, Dong Luo, Erika C. Crouch, Tanya M. Cafarella, James F. Head, Henk P. Haagsman, and Barbara A. Seaton. Journal of Biological Chemistry, 287:26666-26677, 2012.
High-performance scalable molecular dynamics simulations of a polarizable force field based on classical Drude oscillators in NAMD. Wei Jiang, David J. Hardy, James C. Phillips, Alexander D. MacKerell Jr., Klaus Schulten, and Benoît Roux. Journal of Physical Chemistry Letters, 2:87-92, 2011.
Probing a structural model of the nuclear pore complex channel through molecular dynamics. Lingling Miao and Klaus Schulten. Biophysical Journal, 98:1658-1667, 2010.
Flow-induced β-hairpin folding of the glycoprotein Ibα β-switch. Xueqing Zou, Yanxin Liu, Zhongzhou Chen, Gloria Ines Cárdenas-Jirón, and Klaus Schulten. Biophysical Journal, 99:1182-1191, 2010.
Challenges in protein folding simulations. Peter L. Freddolino, Christopher B. Harrison, Yanxin Liu, and Klaus Schulten. Nature Physics, 6:751-758, 2010.
O2-reactivity of flavoproteins: Dynamic access of dioxygen to the active site and role of a H+ relay system in D-amino acid oxidase. Jan Saam, Elena Rosini, Gianluca Molla, Klaus Schulten, Loredano Pollegioni, and Sandro Ghisla. Journal of Biological Chemistry, 285:24439-24446, 2010.
Molecular dynamics simulations suggest that electrostatic funnel directs binding of Tamiflu to influenza N1 neuraminidases. Ly Le, Eric H. Lee, David J. Hardy, Thanh N. Truong, and Klaus Schulten. PLoS Computational Biology, 6:e1000939, 2010. (13 pages).
Limits for reduction of effective focal volume in multiple-beam light microscopy. Anton Arkhipov and Klaus Schulten. Optics Express, 17:2861-2870, 2009.
Transport-related structures and processes of the nuclear pore complex studied through molecular dynamics. Lingling Miao and Klaus Schulten. Structure, 17:449-459, 2009.
Double stranded DNA dissociates into single strands when dragged into a poor solvent. Shuxun Cui, Jin Yu, Ferdinand Kühner, Klaus Schulten, and Hermann E. Gaub. Journal of the American Chemical Society, 129:14710-14716, 2007.
Molecular dynamics simulations of the complete satellite tobacco mosaic virus. Peter L. Freddolino, Anton S. Arkhipov, Steven B. Larson, Alexander McPherson, and Klaus Schulten. Structure, 14:437-449, 2006.
Finding gas diffusion pathways in proteins: Application to O2 and H2 transport in CpI [FeFe]-hydrogenase and the role of packing defects. Jordi Cohen, Kwiseon Kim, Paul King, Michael Seibert, and Klaus Schulten. Structure, 13:1321-1329, 2005.
Binding dynamics of isolated nucleoporin repeat regions to importin-β. Timothy A. Isgro and Klaus Schulten. Structure, 13:1869-1879, 2005.
Conformational model of the Holliday junction transition deduced from molecular dynamics simulations. Jin Yu, Taekjip Ha, and Klaus Schulten. Nucleic Acids Research, 32:6683-6695, 2004.
Genetically engineered gold-binding polypeptides: Structure prediction and molecular dynamics. Rosemary Braun, Mehmet Sarikaya, and Klaus Schulten. Journal of Biomaterials Science, 13:747-758, 2002.
Research Projects
- Kinetic Diffusion
- Oil and Water Split DNA
- Microscopy and Computing Sharpen the Focus
- Spelunking Inside Myoglobin
- Graphics Processors Speed Up Simulations
- Passport for the Cell's Nucleus
- Bringing Oxygen into an Enzyme
- Cryptochromes and Magnetic Sensing
- The Ins and Outs of the Nucleus
- Viruses Up Close
- LOV in Motion
- Molecular Dynamics of STMV
- Gateway to the Nucleus
- Conformational Change of Holliday Junction
- The Nuclear Pore Complex
- Gas Transport Inside Hydrogenase
- Estrogen Receptor Interacting with DNA
- Gold Binding Proteins
- Imaging gas migration pathways inside myoglobin
- Drude Polarizable Force Field
- Molecular Flow Sensor
- Coarse-Grained Molecular Dynamics
- Modeling Biological Processes at Hybrid Resolutions
- Molecular Mechanisms of Human Immunodefficiency Virus (HIV) infection
- MD Simulation of Protein Folding
- Oxygen Migration in DAAO
- Molecular Mechanism of Tamiflu Resistance
- Molecular Basis of Drug Resistance in A/H1N1 Influenza Virus
- Lung Collectins
- DNA Methylation and Hydroxymethylation
- Target-search Mechanism of Protelomerase TelK
- Excitation Dynamics
- ffTK - Force Field Toolkit
- Multilevel Summation Method



