C&S1: Designing Antibodies Targeting Viral Glycoproteins
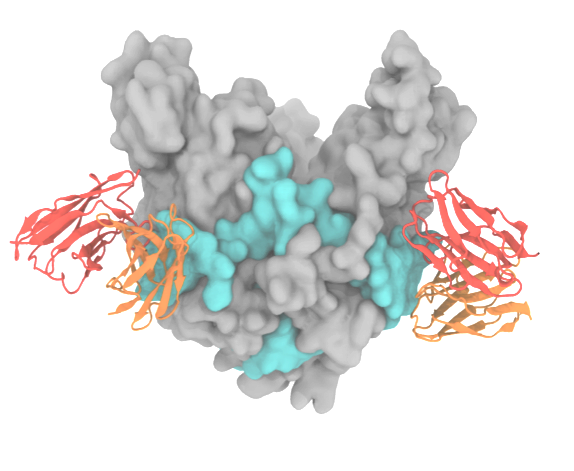
Cell entry is the first stage of the viral infection cycle. Enveloped viruses such as influenza A and Ebola employ glycoproteins (GP), located on the viral surface, to facilitate cell entry and exit. Inhibiting the viral GP by using drugs or antibodies is therefore an effective way to impair the virus's ability to enter the cell. In particular, antibodies that bind with high affinity and specificity are valuable as they can be used for diagnostic and therapeutic purposes. Despite its biomedical importance, developing novel antibodies using solely experimental methods remains a time-consuming and expensive process. Center-developed technologies are employed to improve a computational antibody design method that targets viral GP.
Computational methods had been developed for de novo antibody design. However, these methods could not accurately design antibodies against flexible antigens as they only account for static structures. Therefore, MD simulation performed using NAMD (TRD1) is incorporated in the antibody design protocol to account for the dynamic properties of antigens and antibodies, optimizing the antigen-antibody binding interface. Large computational resources and long turnaround times are the main factors which have hitherto prevented the antibody design community from using MD simulations. The demand for large computational resources is especially evident in our study, noting that hundreds of simulations have to be performed with antibodies in complex with the Ebola GP, a large trimeric structure composed of more than 1,000 residues. To reduce the turnaround time, the simulations will be performed on supercomputers using NAMD (TRD1). NAMD is highly suited for such simulations because of its scalability and the usage of GPU that at least doubles the computing performance. Automated protocols have also been developed for rapid system preparations using VMD (TRD2). The incorporation of NAMD and VMD helps surmount MD simulations as the computational bottleneck in the antibody design protocol.
The study of viruses and antiviral compounds will build upon Center's work on virus capsids (DBP 6) and virus GP. Particularly, the Center has investigated the inhibition of GP of influenza A virus through drugs like Tamiflu. The Center has also collaborated with Seaton who provided crystallographic structures of antiviral peptides to study their inhibitory mechanisms against influenza A virus and other antigens. Recently, the Center has incorporated MD simulation into OptMAVEn, an antibody design framework first developed by Maranas. The improved framework, termed OptMAVEn-MD, has successfully designed antibodies targeting a short and flexible peptide antigen, which were subsequently validated by experiments. The techniques developed in the computational-experimental collaboration are currently being employed in designing novel antibodies against the Ebola virus.
We will work closely with Maranas to continue improving OptMAVEn-MD such that antibodies will be accurately developed to target larger biomolecules such as Ebola GP. The antibodies with high binding affinity will be experimentally characterized using a high-throughput screening assay developed by Tessier. Tessier will also modify the antibodies to increase their solubility and to reduce the propensity to aggregate. As the nature of the project requires close collaboration between the Center, Maranas and Tessier, turnaround time for MD simulations must be minimized. The high performance of NAMD on supercomputers (TRD1) allows hundreds of antibody designs be simulated and tested simultaneously, effectively reducing the turnaround time to less than a week. Remote visualization (TRD2) will be used to communicate simulation results with Maranas and Tessier. The TimeLine feature in VMD (TRD2) will be used to visualize and analyze the key GP-antibody interactions. The antibody design protocol developed in this collaboration can be used to target other viral antigens such as HIV GP and the envelope proteins of Dengue and Zika viruses.



