Resource Statistics Workshop on Computational Biophysics 2026
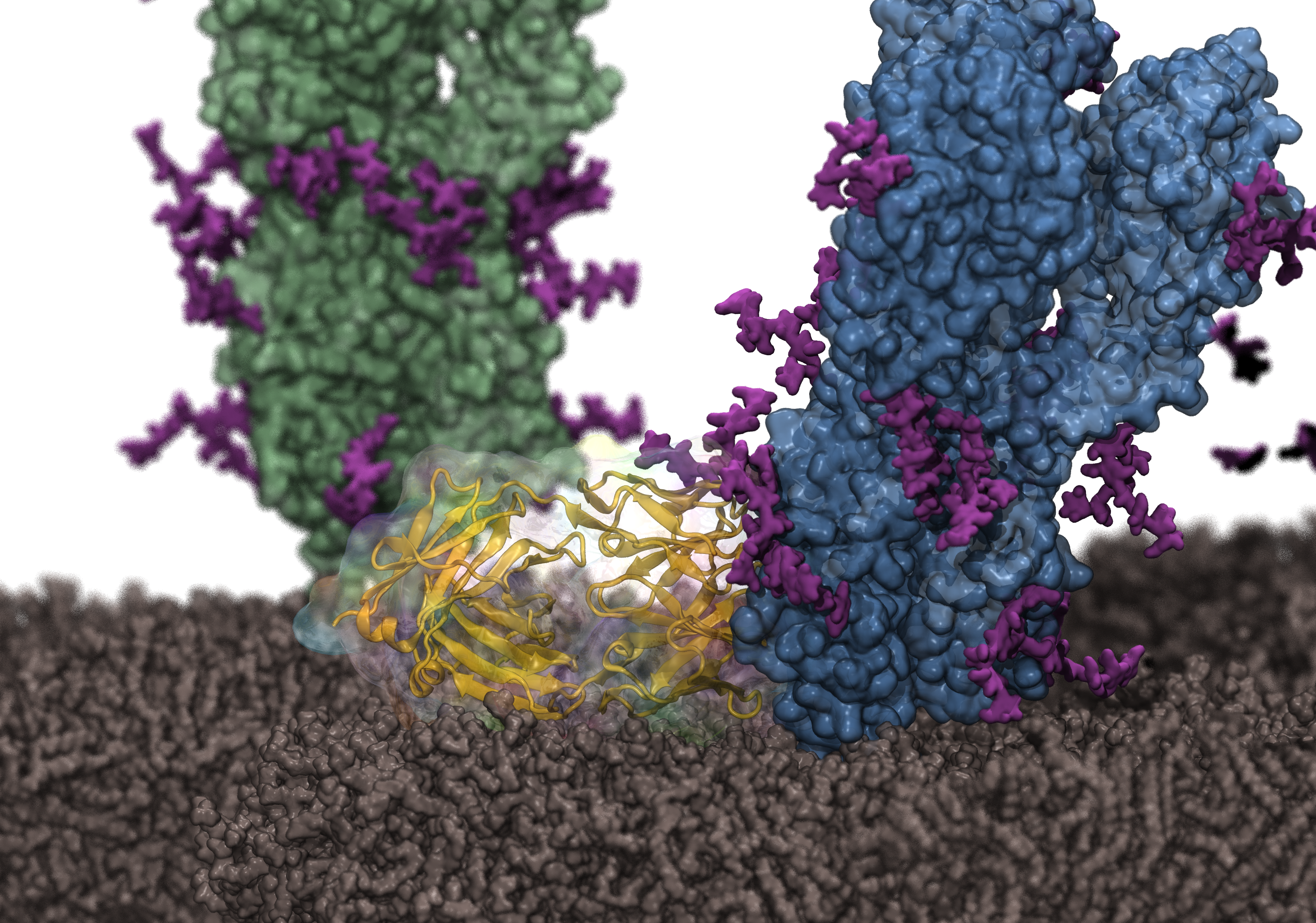
Follow \@tcbguiucInfluenza hemagglutinin (HA) is the most abundant surface antigen of the virus. Since there are 18 subtypes of HA with different antigenicity, there is a strong need to develop broadly neutralizing antibodies targeting multiple subtypes. In collaboration with the Wu Lab at the University of Illinois, Resource researchers reconstructed the structure of the complex between HA and a broadly neutralizing antibody on the viral membrane and simulated it with NAMD. The simulation results revealed membrane interactions for the antibody, which were then confirmed by mutagenesis experiments. The results are made recently available as a cover article in Structure and provide a foundation for rational design of more effective antibodies.
The Future of Biomolecular Modeling
A 2015 TCBG Symposium brought together scientists from across the Midwest to brainstorm about what's on the horizon for computational modeling. See a summary of what these experts foresee.
Read more
The Annals of MDFF
MDFF is a computational method that yields structures of massive biomolecular assemblies at atomic detail, using hybrid experimental data. Now Illinois scientists are applying MDFF to fascinating systems like the ribosome and HIV. By Lisa Pollack.
Read more
Announcements
Seminars
Resource Usage
Remembering Klaus Schulten
Recent Publications All Publications
- Probing Solution Dynamics of Tissue Factor Using Molecular Dynamics Simulations Guided by NMR Chemical Shifts. J. Phys. Chem. B, 2026.
- Elucidating the binding and metabolic interactions of sunitinib and sorafenib with cytochrome P450s CYP2U1 and CYP2D6. Mol. Pharmacol., 108(4):100114. 2026.
- LetA defines a structurally distinct transporter family. Nature, 651(8107): 1097–1106. 2026.
- Single-Molecule Electron Transport in Peptoids. J. Phys. Chem. B, 130(11):3054–3064. 2026.
- Cholesterol efflux protein, ABCA1, supports anticancer functions of myeloid immune cells. Sci. Adv., 12(1):eadx5490. 2026.
- Thermodynamic and Kinetic Analysis of Molecular Conformational Dynamics in a Riemannian Framework. J. Phys. Chem. A, 130(5):1220–1232. 2026.
- Simulating Gas Permeation Through the Central Pore of AQP5. Adv Exp Med Biol., 1498:105–113. 2026.
Highly Cited
Lipid bilayer pressure profiles and mechanosensitive channel gating. Biophysical Journal, 86:3496-3509, 2004.
Click here for other highly cited papers
Click here for other highly cited papers